Introduction
This guide explains how to publish your biodiversity data to GBIF and the world using the Canadensys repository. It is not the only method you can use to publish your data, but we think it is presently the most convenient one for Canadian collections and organizations. Our repository is powered by the GBIF Integrated Publishing Toolkit (IPT) and maintained by us, which allows you to upload, standardize, publish and register your data in 7 steps, without the hassle of installing and maintaining a program. The data are published in your organization’s name and it is free.
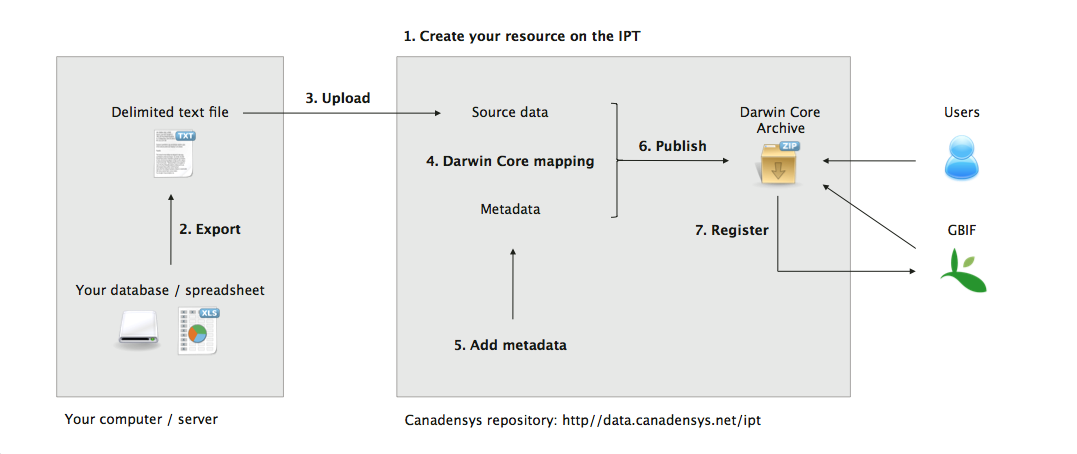
For information about the different classes of datasets, see the following GBIF guides:
- Data quality requirements to publishing occurrence data
- Data quality requirements to publishing checklists
- Data quality requirements to publishing metadata
- Data quality requirements to publishing sampling-event data
Conditions
We care about data and we just want to make sure you do too. In order to publish your data using the Canadensys repository you should meet the following criteria:
- You are associated with a Canadian collection or organization.
- You are publishing a specimen or observation dataset, a taxonomic checklist, a sampling-event dataset or simply metadata (in other words, one of the 4 types of datasets supported by the IPT).
- You hold the rights to publish the data.
- You are willing to maintain the dataset and improve its quality where possible.
- You are willing to provide sufficient metadata, so users can learn what the dataset is about.
- You are publishing the data under an open license, so others can really use them. We strongly recommend publishing under CC0.
1. Create your resource on the IPT
The Canadensys repository is powered by the GBIF Integrated Publishing Toolkit (IPT), an open source web application developed by GBIF and customized by Canadensys to be fully integrated into our website. We use it to publish and register all our datasets. To be able to create and manage your own dataset (called a “resource”), you will need a user account. Just contact us to create one for you.
Once you have your account, login at the top of this page. Click on the tab Manage resources to get access to your dashboard. It will display all the datasets you are managing and will be empty at first. You can create a new resource at the bottom of the page. Follow the IPT manual for more detailed instructions.
Warning: please use the following lowercase format for the shortname of your resource: yourcollectioncode-datasettype (e.g. acad-specimens or wildlife-sightings-observations). This name is used to uniquely identify and access your resource and cannot be modified subsequently! For testing purposes, please use yourcollectioncode-test (e.g. ubc-test).
Once you have created your resource, you will see an empty resource overview page.
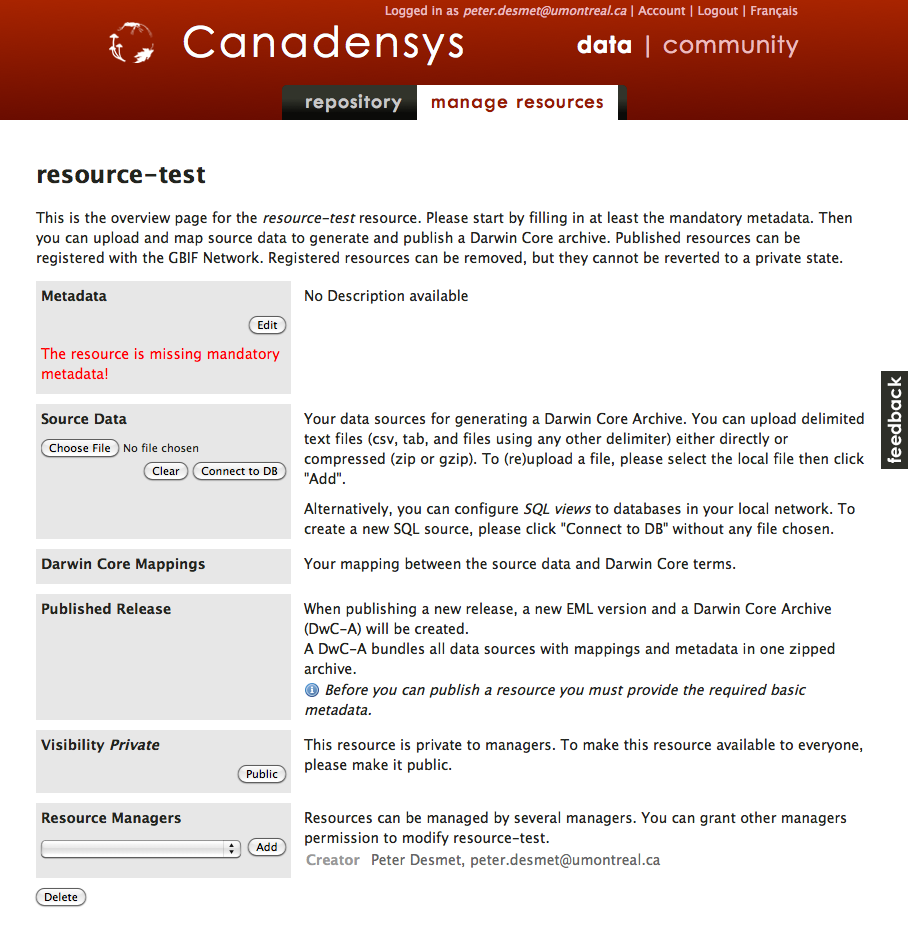
2. Export
The easiest way to get your data into the IPT is to first export them from your database as a delimited text file (e.g. .txt, .tab, .csv). Most databases offer the option to do so. Use the UTF-8 character encoding for your export (and not ASCII, Macintosh or Windows ANSI) to avoid misinterpretations of accented characters (e.g. é, à, ü, î). If you have the choice, include a header line in your export (a first line with the field names), as it will be helpful later.
3. Upload
Uploading your source file to the IPT is easy: go to your resource overview page > Source Data and click Choose File. You might want to compress/zip your source file first to improve the upload speed of large files. The IPT will unzip them automatically once received. Follow the IPT manual for more detailed instructions (including the option to use multiple source files or to upload via a direct database connection).
Once your source file has been uploaded correctly, a source file detail page will be shown (see an example screenshot in the IPT manual), displaying how the IPT has interpreted your file (number of columns, rows, header rows, character encoding, delimiters, etc.). Click the preview button to verify everything is correct, then click save.
4. Darwin Core Mapping
Biodiversity data are published in the Darwin Core standard. It includes a list of defined terms and allows your data to be understood and used by anyone. It also allows an aggregator like GBIF to combine your data with other data, like they do on their portal.
Darwin Core mapping is the process of linking the fields in your source file with the appropriate Darwin Core terms. It is the most challenging step in publishing your data for two reasons: 1) the list of Darwin Core terms can be overwhelming, so it might be difficult to select the ones that are appropriate for your dataset, and 2) the IPT currently only allows one-to-one mapping of fields, so the ease of mapping will depend on your database structure and on the feasibility of exporting as close to Darwin Core as possible.
This is why we are here to help! Contact us to arrange a phone or online call to guide you through the steps, review your mapping, suggest terms and help you repeat steps 2-4 until the mapping is just right.
You can find more information regarding Darwin Core mapping in the IPT manual (including core types, extensions, automapping, default values, value translation, etc.) and in the introduction to Darwin Core on our website.
5. Add metadata
If data are LEGO bricks, then metadata are the shiny box and instructions. They enable users to discover your dataset and assess its relevance for their particular needs, so it pays off investing some time providing them.
Go to your resource overview page > Metadata and click Edit to open the metadata editor. Contact us to register your institution (if not done so already) to GBIF, in order to be able to link your resource with your institution in the metadata.
Any information you provide here will be visible on the resource homepage and bundled together with your data when you publish. Metadata are expressed in the GBIF EML Profile standard and can also be downloaded as a Rich Text Format (RTF) file. The latter can serve as a draft manuscript describing the dataset (a “Data Paper“), which can be submitted for peer-review to a Pensoft open-access journal such as the Biodiversity Data Journal, Phytokeys, Zookeys, Biorisk, Neobiota or Nature Conservation.
Follow the IPT manual for detailed instructions about the metadata editor and use one of the currently published datasets as an example (e.g. collection example, checklist example). See our website for more information regarding metadata.
6. Publish
At this stage, you are all set to publish! Go to your resource overview page > Published Versions and click Publish. The IPT will now generate your data as Darwin Core, combine it with the metadata and package it as a standardized zip-file called a “Darwin Core Archive“.
If you wish to assign a DOI to your dataset, you need to click on the ‘Reserve’ button, then change the visibility of your dataset to ‘Public’ before hitting the ‘Publish’ button. The DOI is registered directly with DataCite and a link will be added in the metadata of your ressource. For more information about the DOI assignation steps, see the IPT DOI Workflows. See the IPT manual for more details about the general publication process.
Back on the resource overview page > Published Versions, you can see the details of your first published dataset, including the publication date and the version number. Since your dataset is published privately, the only thing left to do is to click Visibility > Public (see the IPT manual) to make it available to everyone. Warning: please do not do this for your test dataset.
Congratulations, you just published your dataset to the world! It is now listed on the repository homepage and you can share and link to it via: https://data.canadensys.net/ipt/resource?r=dataset-shortname. This would be a good time to notify any regional or thematic network you are involved in, such as VertNet, the Consortium of Northeastern Herbaria or the Entomological Society of Canada.
Your published dataset is a static snapshot of your data and will not change until you upload an updated source file and click publish again. This procedure has the advantage that your dataset is always available, does not require a live internet connection to your database and can be easily shared (e.g. you can email the Darwin Core Archive to a colleague). It also allows you to control the publication process more precisely: version 1, version 2, etc. and users are informed of how recent the data are and the differences between versions (addition of data, correction of errors, etc).
7. Register with GBIF
Even though your dataset is now available to everyone, it might be difficult for users to discover it. This is why we recommend registering your dataset with the Global Biodiversity Information Facility (GBIF). It allows your data to become available to an international audience via the GBIF portal and it ensures full attribution is given to your institution. By registering, you agree with the GBIF Data Publisher Agreement.
On the resource overview page, click on Visibility > Register (see the IPT manual) to register your dataset with the GBIF registry. It will allow GBIF to index your resource on their portal, where it can be easily accessed by everyone.
Citation
As all documents on this website, this guide is published under CC-BY. The preferred citation is:
Desmet, P. & C. Sinou. 2012. 7-step guide to data publication. Canadensys. https://canadensys.net/publish/7-step-guide/